“Once sequenced, a cancer tumor can have thousands of genetic mutations. But the challenge is distinguishing the mutations that contribute to tumor growth (drivers) from the neutral mutations (passengers).
Currently this interpretation of genetic mutations is being done manually. This is a very time-consuming task where a clinical pathologist has to manually review and classify every single genetic mutation based on evidence from text-based clinical literature.We need your help to develop a Machine Learning algorithm that, using this knowledge base as a baseline, automatically classifies genetic variations.”
~~Kaggle
_For implementation please follow the link : _https://github.com/vedanshsharma/Personalized-Cancer-Diagnosis
Introduction
The task of identifying the type of variation of the gene is typically a three step process. Our task is to use a model to automate the third step which is the most time consuming step for a molecular pathologist. This involves analyzing the evidence related to each of the variations to classify them. More formally our problem statement is -
Classify the given genetic variations/mutations based on evidence from text-based clinical literature.
Source: https://www.kaggle.com/c/msk-redefining-cancer-treatment/discussion/35336#198462
Business objectives and constraints.
- No low-latency requirement. Since we don`t have a strict latency requirement we can train complex models as long as we our model is taking reasonable amount of time.
- **Interpretability is important. **A doctor might want to cross check the results and for this he might require the algorithm to explain itself. For this interpretability should be high.
- **Errors can be very costly. **Since this is a matter of life and death , we need a model with high accuracy.
- Probability of a data-point belonging to each class is needed. This will let the doctor decide at the end to which class the variation belongs to. In case of equiprobable classes, the doctor will be able to decide by performing tests, rather than the model returning a single class.
Machine Learning Problem Formulation
Data
- We have two data files: one contains the information about the genetic mutations and the other contains the clinical evidence (text) that human experts/pathologists use to classify the genetic mutations.
- Both these data files are have a common column called ID
- Data file’s information:
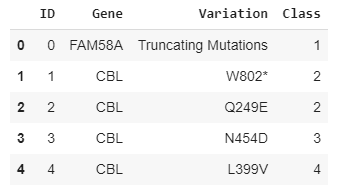
- training_variants (ID , Gene, Variations, Class)
- training_text (ID, Text)

Mapping the real-world problem to an ML problem
There are nine different classes a genetic variation/mutation can be classified into. This implies that our problem is a **multi class classification **problem. The performance metrics that we will be using are -
- Multi class log-loss, Since we want to output probabilities that a given variable x_i belong to classes 1–9.
- Confusion matrix
So our final objective is-
Objective: Predict the probability of each data-point belonging to each of the nine classes.
And our final constraints are-
- Interpretability ,models like naive bayes , linear svm or DTs can be trained as they have high interpretability. We should probably avoid using models like RBF svm.
- Class probabilities are needed.
- Penalize the errors in class probabilites => Metric is Log-loss.
- No Latency constraints.
Train, CV and Test Datasets
Our data is not temporal in nature i.e. it doesn`t change with time. Hence we will be randomly splitting our data set into train Cv and test set with sizes being 64% (80 % of 80), 16 %(20% of 80) and 20% respectively.We will split the data in such a way that the distribution of class label remain conserved.
Exploratory Data Analysis
Text Preprocessing
We will be doing some basic text preprocessing which includes-
- Removing stop words.
- replacing every special character with space.
- replacing multiple spaces with single space.
- converting all the chars into lower-case.
There are few null values present in text.
#cancer #machine-learning #random-forest #data analysis
